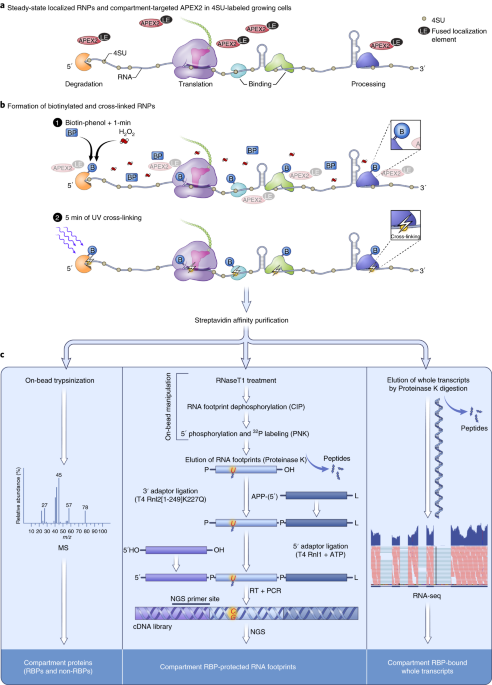
Proximity-CLIP provides a snapshot of protein-occupied RNA elements in subcellular compartments | Nature Methods

Screening strategies for identifying RNA- and ribonucleoprotein-targeted compounds: Trends in Pharmacological Sciences
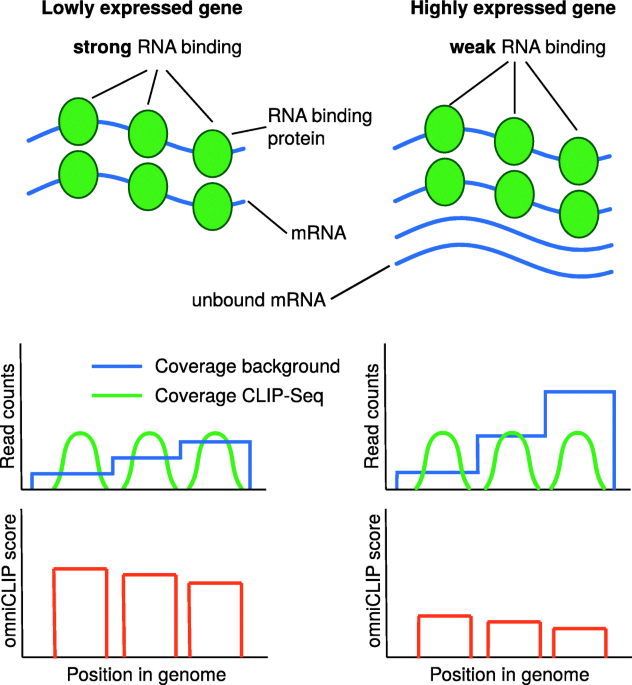
omniCLIP: probabilistic identification of protein-RNA interactions from CLIP-seq data | Genome Biology | Full Text
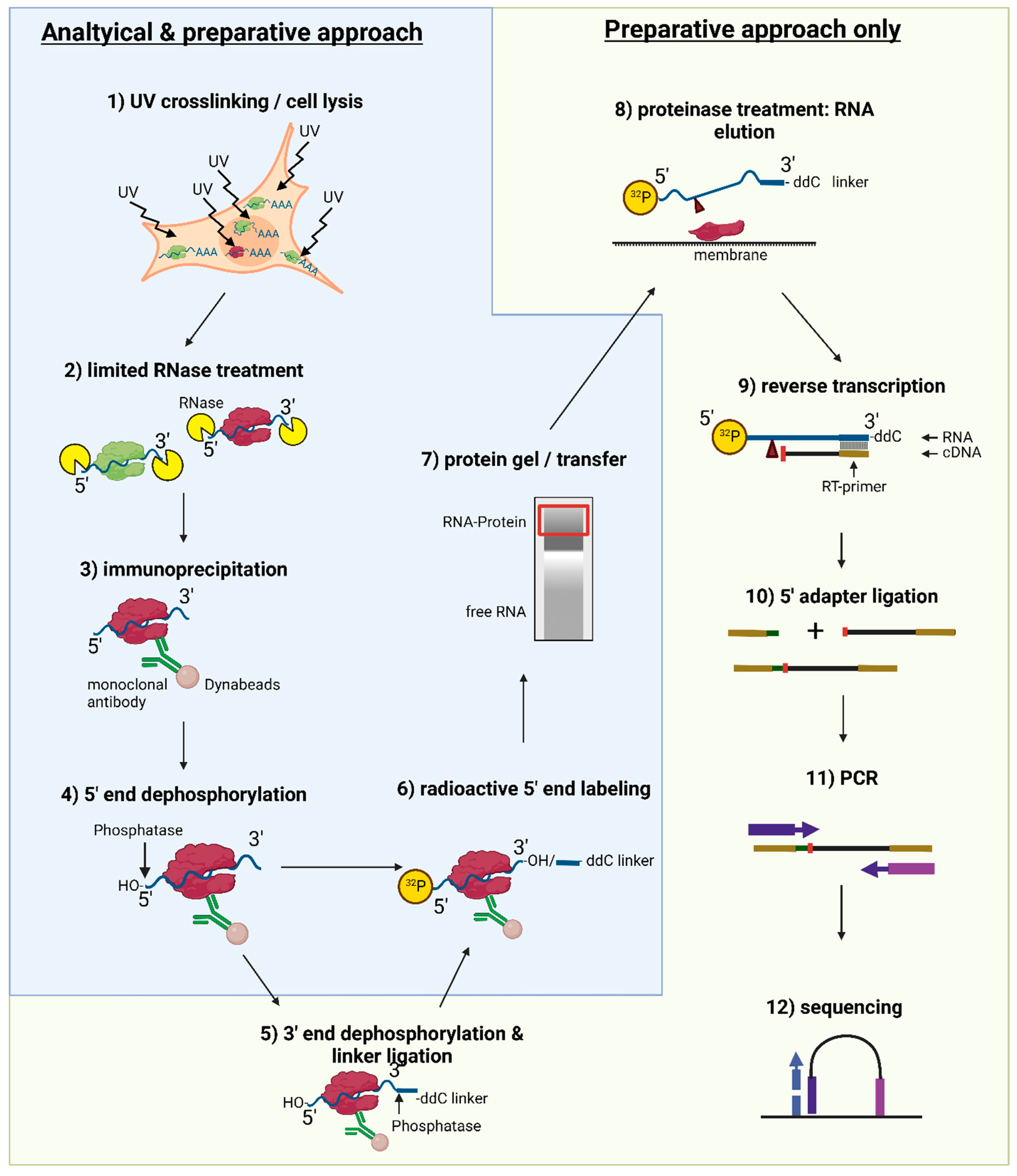
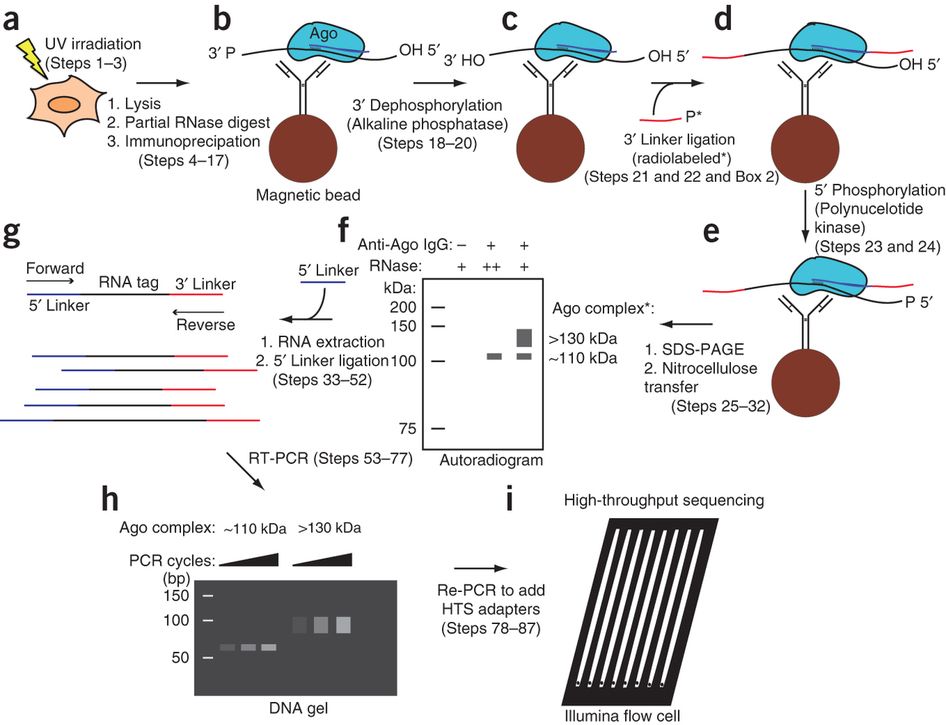
MPs | Free Full-Text | Studying miRNA–mRNA Interactions: An Optimized CLIP-Protocol for Endogenous Ago2-Protein
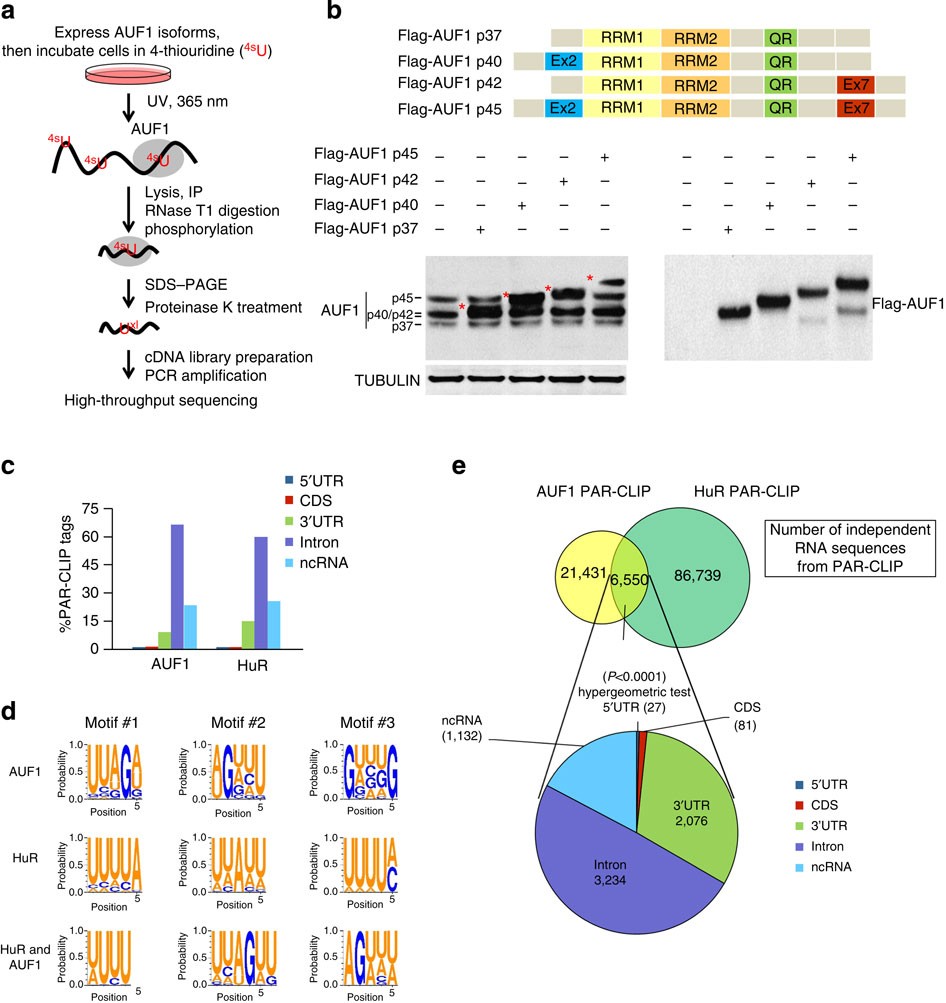
PAR-CLIP analysis uncovers AUF1 impact on target RNA fate and genome integrity | Nature Communications

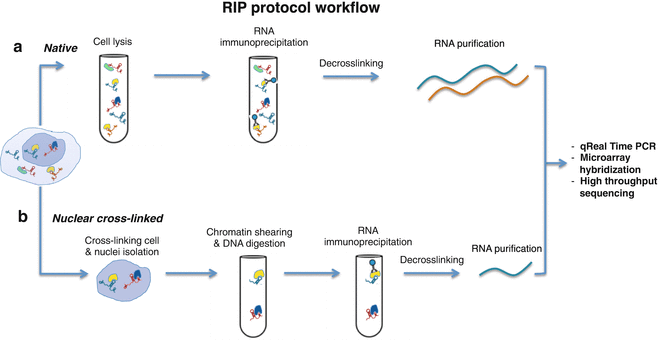
Crosslinking Immunoprecipitation and qPCR (CLIP-qPCR) Analysis to Map Interactions of Long Noncoding RNAs with Canonical and Non-canonical RNA-Binding Proteins | SpringerLink

Analysis of CLIP and iCLIP methods for nucleotide-resolution studies of protein-RNA interactions | Genome Biology | Full Text

Denaturing cross-linking immunoprecipitation to identify footprints for RNA-binding proteins - ScienceDirect
Analysis of the nucleocytoplasmic shuttling RNA-binding protein HNRNPU using optimized HITS-CLIP method | PLOS ONE
CLIP variants for studying RBP-RNA interactions (ii). (A) Simplified... | Download Scientific Diagram





-1.jpg)